Die Forschungsarbeiten des Virologischen Instituts konzentrieren sich auf ein wichtiges humanpathogenes Virus, das humane Cytomegalovirus (HCMV). Dabei handelt es sich um ein weit verbreitetes Herpesvirus, das bei etwa 40% der mitteleuropäischen Bevölkerung Infektionen hervorruft. In vielen Fällen verlaufen diese Infektionen ohne eine ausgeprägte Symptomatik. Das Virus persistiert jedoch in einer latenten Form im Körper des Infizierten: es kann im Lauf des Lebens reaktivieren und erneut eine Krankheitssymptomatik hervorrufen. Dies kommt sehr häufig bei eingeschränkter Immunfunktion vor, so dass HCMV als einer der wichtigsten Krankheitserreger bei immunsupprimierten Patienten (z.B. Transplantationspatienten) gilt. Darüber hinaus kann dieses Virus während der Schwangerschaft eine Infektion des ungeborenen Kindes hervorrufen, die dauerhafte und z.T. schwerwiegende Schädigungen beim Kind verursacht. In unseren Forschungsarbeiten charakterisieren wir molekulare Details des Viruseintritts in die Zielzelle (AG Sinzger →), der viralen Genexpression (AG Stamminger →) sowie der Virusmorphogenese (AG von Einem →), um neue antivirale Therapieansätze und Medikamente zu entwickeln. Zusätzlich beschäftigen wir uns mit offenen Fragen der viralen Pathogenese sowie der Analyse von viralen Resistenzmechanismen (AG Michel →).
Research group of Prof. Dr. Thomas Stamminger
HCMV regulatory proteins and innate immunity
Human cytomegalovirus (HCMV) remains an important cause of mortality and morbidity in immunocompromised patients such as transplant recipients on immunosuppressive therapy. Furthermore, HCMV is the most common cause of intrauterine infection which can lead to sensorineural hearing loss and mental retardation. Although progress in the diagnosis of HCMV infections has led to improved therapeutic strategies, treatment of HCMV induced disease suffers from the toxicity of the currently available antiviral drugs. In addition, in patients where longer therapy is necessary the emergence of drug-resistant viral strains is frequent. Therefore, the development of novel antiviral drugs is urgently required in order to improve the therapy of HCMV infections. Thus, one important aim of our laboratory is the characterization of molecular events that can be used as targets for new antiviral strategies.

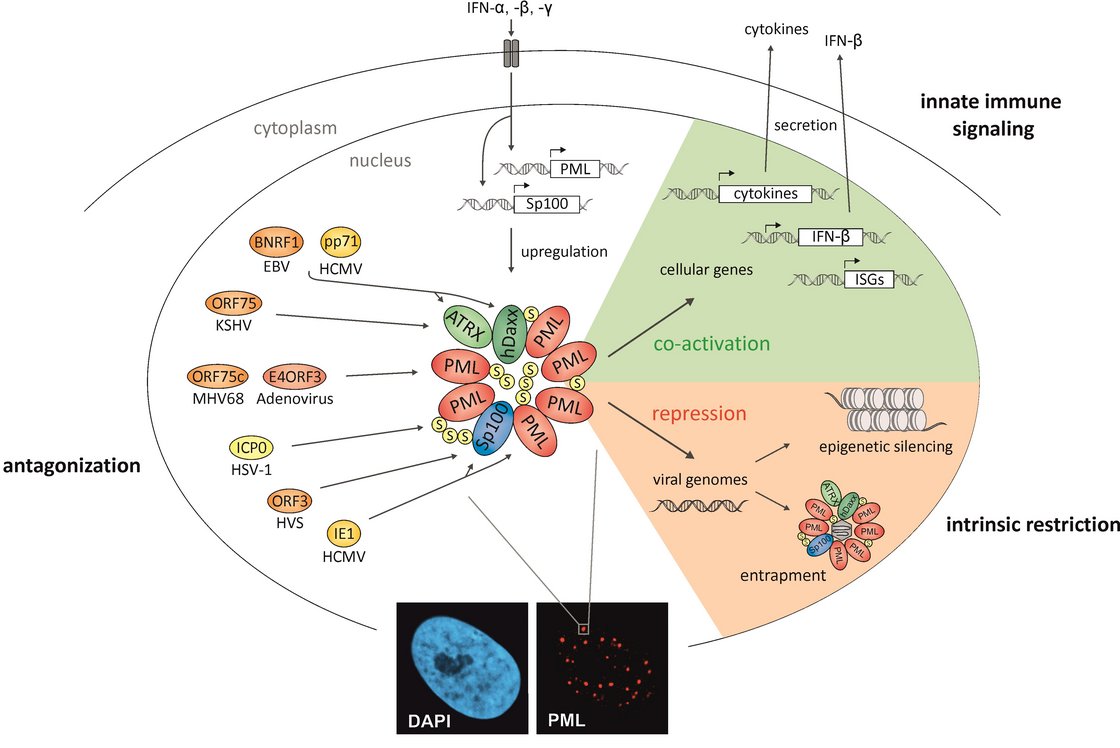
PML nuclear bodies: a cellular structure that mediates intrinsic immunity against viruses. Microbial infections are not only controlled by innate and adaptive immune mechanisms but also by cellular restriction factors which give cells the capacity to resist pathogens. Unlike the innate and adaptive part of the immune system, that require pathogen-induced signaling cascades in order to be switched on, these so-called intrinsic immune mechanisms are mediated by cellular proteins that are constitutively expressed and active before a pathogen enters the cell, thus serving as a front-line defense. Our laboratory investigates whether a subnuclear structure, termed PML nuclear bodies (PML-NBs), contributes to the resistance of cells against herpesvirus infections. PML-NBs are dot-like structures of the cell nucleus, that are defined by the distinct accumulation of specific cellular proteins like PML, hDaxx, Sp100 and ATRX (Fig. 1). During the last ten years our laboratory could show, that these proteins act as cellular restriction factors by inducing a silencing of viral gene expression. Moreover, our research revealed that specific viral proteins (pp71 and IE1 of HCMV) are able to antagonize this cellular silencing mechanism (Fig. 1). This establishes a delicate balance between cellular defense and viral antagonism which determines whether the herpesvirus enters the productive cycle or viral gene expression is silenced.
Fig. 1: PML-NBs: a cellular structure that mediates both intrinsic immunity against viruses and acts as a coactivator of the interferon response. Viral effector proteins that are known to antagonize PML-NBs are shown in the left part of the figure.
Molecular mechanism of PML-NB disruption. During the last two years we could delineate the molecular mechanism how IE1 antagonizes PML-NBs. As evident from the crystal structure of IE1 that we recently solved in collaboration with Prof. Y. Muller and Prof. H. Sticht, this protein directly interacts with the PML coiled-coil domain via its globular core region and disrupts NB foci by inducing a loss of PML SUMOylation. We were able to demonstrate that IE1 acts via abrogating the de novo SUMOylation of PML. In order to overcome reversible SUMOylation dynamics, we made use of a cell-based assay that combines inducible IE1 expression with a SUMO mutant resistant to SUMO proteases. Interestingly, we observed that IE1 expression did not affect preSUMOylated PML, however, it clearly prevented de novo SUMO conjugation. Consistent results were obtained by in vitro SUMOylation assays demonstrating that IE1 alone is sufficient for this effect. Furthermore, IE1 acts in a selective manner since K160 was identified as the main target lysine. This is strengthened by the fact that IE1 also prevents As2O3-mediated hyper-SUMOylation of K160 thereby blocking PML degradation. Since IE1 did not interfere with coiled-coil mediated PML dimerization we propose that IE1 either affects PML autoSUMOylation by directly abrogating PML E3 ligase function or by preventing the access to SUMO sites. Thus, our data suggest a novel mechanism how a viral protein counteracts a cellular restriction factor by selectively preventing the de novo SUMOylation at specific lysine residues without affecting global protein SUMOylation.
PML nuclear bodies as coregulatory structures of innate immune responses. Studies performed during the last years provided evidence for an extended cross-talk between intrinsic and innate immune mechanisms in relation to PML-NBs. For instance, specific PML-NB factors such as PML and Sp100 are known to be induced by interferons (IFN) and the overall size and number of PML-NBs also increases upon IFN treatment of cells. This indicates that the restrictive function of PML-NBs towards viral infections can be enhanced via IFNs. However, recent evidence now suggests that PML itself acts as an important co-regulatory factor during the induction of interferon stimulated genes (ISGs) (Fig. 1). For instance, we observed in collaboration with PD. Dr. P. Hemmerich (Jena) that the PML protein is able to promote interferon-g (IFN-g) induced MHC class II gene expression. However, the activities of PML as a coactivator of the IFN response are not confined to type II IFNs but also extend to type I IFN-regulated genes. This was observed in our recent study that detected a significantly reduced induction of specific ISGs after type I IFN treatment of cells exhibiting a depletion of the endogenous PML protein. The molecular mechanism by which PML stimulates the induction of type I IFN-regulated genes is not completely clear so far, but may involve the PML-NB-mediated stabilization of transcriptional complexes that control ISG expression. Consequently, as we were able to demonstrate for the IE1 protein of HCMV, viral proteins that modulate PML-NBs may not only antagonize intrinsic antiviral defense but also innate immune responses.
Signaling of HCMV-encoded G protein-coupled receptors. G protein-coupled receptors (GPCRs) are key regulators of numerous cellular processes. Thus, virally encoded GPCRs illustrate an effective means to bypass the immune system, modulate cellular functions and redirect cellular signaling networks. Human cytomegalovirus (HCMV) encodes four GPCR homologues, termed pUS27, pUS28, pUL33, and pUL78. Whereas pUS28 and pUL33 constitutively activate multiple signaling cascades, the functions of pUS27 and pUL78 are not fully understood, yet. To determine whether pUS27 or pUL78 display any signaling properties, we performed luciferase reporter assays in 293T cells transduced with CREB-, NFAT and NF-κB-specific reporter constructs. Our experiments demonstrate for the first time that pUS27 activates NF-κB dependent gene expression. Intriguingly, it turned out that NF-κB activation differed significantly depending on whether N- or C-terminally tagged versions of pUS27 were applied: while transfection of N-terminally tagged pUS27 did not activate NF-κB, the expression of C-terminally tagged versions strongly induced NF-κB signaling. Disruption of a putative PDZ domain binding motif by adding one serine to the C-terminus of pUS27 induced high NF-κB signaling suggesting that pUS27 may be negatively regulated via a PDZ domain protein. Bioinformatic analysis revealed the existence of four putative TRAF binding motifs within the C-terminus of pUS27. Interestingly, mutation of a predicted TRAF6 binding motif led to a complete loss of NF-κB signaling. To answer the question whether the C-terminal region of pUS27 alone is essential and sufficient for NF-κB activation, we generated chimeric proteins with either CD8 or GFP. These chimeras strongly activated NF-κB signaling independent of their localization at either the cell surface or endosomes, respectively. This indicates that the subcellular localization of the pUS27 cytoplasmic domain is not critical for NF-κB activation. Moreover, we could show that pUS27 specifically induced the canonical NF-κB pathway through TRAF6 by using either ACHP, an inhibitor of IKKβ phosphorylation, or a dominant negative IκBα. Taken together, our data reveal a novel and highly complex signaling capability of the HCMV GPCR pUS27.
The UL69 of human cytomegalovirus: a regulator of viral mRNA export that is controlled by distinct post-translational modifications. The open reading frame UL69 of human cytomegalovirus encodes the multi-functional, regulatory protein pUL69 which acts as a viral mRNA export factor via its nucleocytoplasmic shuttling activity and via recruitment of the cellular mRNA export machinery by interacting with the cellular DExD/H-box RNA helicase UAP56. We reported previously, that subcellular localization and mRNA export activity of pUL69 are modulated via phosphorylation by the viral kinase pUL97. In recent studies were able to obtain evidence that pUL69 is subject to additional posttranslational modifications like arginine methylation. First, we demonstrated a specific immunoprecipitation of full-length pUL69 as well as pUL69aa1-146 by a mono/dimethylarginine-specific antibody. Second, we observed a specific electrophoretic mobility shift upon overexpression of the catalytically active protein arginine methyltransferase 6 (PRMT6). Third, a direct interaction of pUL69 and PRMT6 was confirmed by yeast two-hybrid and coimmunoprecipitation analyses. We mapped the PRMT6 interaction motif to the pUL69 N terminus and identified critical amino acids within the arginine-rich R1 box of pUL69 that were crucial for PRMT6 and/or UAP56 recruitment. In order to test the impact of putative methylation substrates on the functions of pUL69, we constructed various pUL69 derivatives harboring arginine-to-alanine substitutions and tested them for RNA export activity. Thus, we were able to discriminate between arginines within the R1 box of pUL69 that were crucial for UAP56/PRMT6-interaction and/or mRNA export activity. Remarkably, nuclear magnetic resonance (NMR) analyses revealed the same α-helical structures for pUL69 sequences encoding either the wild type R1/R2 boxes or a UAP56/PRMT6 binding-deficient derivative, thereby excluding the possibility that R/A amino acid substitutions within R1 affected the secondary structure of pUL69. We therefore conclude that the pUL69 N terminus is methylated by PRMT6 and that this critically affects the functions of pUL69 for efficient mRNA export and replication of human cytomegalovirus. Furthermore, we set out to identify phosphorylation sites within the functionally important N-terminus of pUL69 and characterized the in vivo importance for HCMV-replication. In silico analyses predicted the existence of 13 serines and 3 threonines as putative phosphorylation sites within the functionally important N-terminal 140 amino acids of pUL69. MS-based phosphosite mapping of immunopurified FLAG-pUL69 identified S18, S51 and T52 as highly probable phosphorylation sites in transfected HEK293T cells. CoIP analyses revealed that combinatorial exchange of S13+15+16+18 to alanine abolished pUL69´s capacity to interact with UAP56. We therefore hypothesize that cellular kinases phosphorylate this serine stretch upstream of alpha-helix1 and thereby determine the recruitment of UAP56-recruitment by pUL69. By comparing the expression profiles of pUL69-mutants that carry individual or combinatorial substitutions of putative phosphosites via Phos-tag-SDS-PAGE, we provide strong evidence that several serines were phosphorylated when pUL97 wildtype but not when its catalytically inactive derivative pUL97-K355M were coexpressed. Since pS46 and pS49 within alpha-helix2 are both preceding a proline, we speculated that they might be substrate for Pin1-mediated peptidyl-prolyl cis-trans-isomerization and confirmed a complex formation of pUL69 and Pin1 by CoIP experiments. NMR-experiments to define the impact of phosphorylation on the structure of pUL69 are in progress. In accordance with our biochemical in vitro studies, preliminary multistep growth-curve analyses revealed a severe growth defect of recombinant HB15-derived viruses that carry S/T to A-substitutions within the pUL69 N-terminus. We therefore identified phosphosites of pUL69 which are required for UAP56- or Pin1-recruitement, respectively, and confirmed their in vivo importance for HCMV replication.
Collaborations
- Prof. Dr. Manfred Marschall, Institute of Clinical and Molecular Virology, University of Erlangen-Nürnberg, Germany
- Prof. Dr. Armin Ensser, Institute of Clinical and Molecular Virology, University of Erlangen-Nürnberg, Germany
- Prof. Dr. Thomas Gramberg, Institute of Clinical and Molecular Virology, University of Erlangen-Nürnberg, Germany
- Prof. Dr. Yves Muller, Division of Biotechnology, Department of Biology, University of Erlangen-Nürnberg, Germany
- Prof. Dr. Heinrich Sticht, Professorship of Bioinformatics, Institute of Biochemistry – Emil-Fischer-Center, University of Erlangen-Nürnberg, Germany
- Prof. Dr. Alexander Steinkasserer, Department of Immune Modulation at the Department of Dermatology, Universitätsklinikum Erlangen, Germany
- PD Dr. Christian Heim, Department of Cardiac Surgery, Universitätsklinikum Erlangen, Germany
- Prof. Dr. Uwe Sonnewald, Division of Biochemistry, Department Biologie, University of Erlangen-Nürnberg, Germany
- PD Dr. Peter Hemmerich, Leibnitz-Institut für Altersforschung - Fritz Lipmann-Institut e.V., Jena, Germany
- Prof. Dr. Pierre Szepetowski, Institut de Neurobiologie de la Mediterranee, Marseille, France
- Prof. Dr. Patrick Hearing, Dept. Molecular Genetics and Microbiology, School of Medicine, Stony Brook University, New York, USA
- Dr. Peter Lischka, AiCuris GmbH & Co KG, Wuppertal, Germany
- Prof. Dr. Jin-Hyun Ahn, Department of Molecular Cell Biology, Sungkyunkwan University School of Medicine, Suwon, Republic of Korea
- Prof. Dr. J. Sinclair, Addenbrooke‘s Hospital, Cambridge, UK
- Prof. Dr. Torgils Fossen, University of Bergen, Norway
- Prof. Dr. Dana Wolf, Hadassah-Hebrew University Medical Center, Virology Divivion, Jerusalem, Israel
Funding
- Deutsche Forschungsgemeinschaft, SFB 796, „Reprogramming of host cells by microbial effectors", B3 (until Dec. 2017), since 2018 STA 357/7-1
- Interdisziplinäres Zentrum für Klinische Forschung Erlangen (IZKF), project A71
- Wilhelm Sander Stiftung Nr. 2016.087.1
Selected References:
- Reuter, N., Schilling, E.M., Scherer, M., Müller, R., Stamminger, T. (2017). The ND10 component promyelocytic leukemia protein acts as an E3 ligase for SUMOylation of the major immediate early protein IE1 of human cytomegalovirus. J. Virol. 91, e02335-16.
- Schilling, E.M., Scherer, M., Reuter, N., Schweininger, J., Muller, Y.A., Stamminger, T. (2017). The human cytomegalovirus IE1 protein antagonizes PML nuclear body mediated intrinsic immunity via the inhibition of PML de novo SUMOylation. J. Virol. 91, e02049-16.
- Lamm, C.E., Scherer, M., Reuter, N., Amin, B., Stamminger, T., Sonnewald, U. (2016). Human promyelocytic leukemia protein is targeted to distinct domains in plant nuclei and colocalizes with nucleolar constitutents in a SUMO-dependent manner. FEBS Open Bio doi: 10.1002/2211-5463.12134.
- Kim, Y.J., Kim, E.T., Lee, M.K., Kwon, K.M., Kim, K.I., Stamminger, T., Ahn, J.H. (2016). Consecutive inhibition of ISG15 expression and ISGylation by cytomegalovirus regulators. PLoS Pathog. 12(8):e1005850.
- Cloarec, R., Bauer, S., Luche, H., Buhler, E., Pallesi-Pocachard, E., Salmi, M., Courtens, S., Massacrier, A., Grenot, P., Teissier, N., Watrin, F., Schaller, F., Adle-Biassette, H., Gressens, P., Malissens, M., Stamminger, T., Streblow, D.N., Bruneau, N., Szepetowski, P. (2016) PLoS One 11(7): e0160176.
- Zheng, Y., Stamminger, T., Hearing, P. (2016). E2F/Rb family of proteins mediate interferon induced repression of adenovirus immediate early transcription to promote persistent viral replication. PLoS Pathog. 12(1): e1005415.
- Kahle, T., Volkmann, B., Eissmann, K., Herrmann, A., Schmitt, S., Wittmann, S., Merkel, L., Reuter, N., Stamminger, T., Gramberg, T. (2015). TRIM19/PML restricts HIV infection in a cell type-dependent manner. Viruses 8(1). Doi: 10.3390/v8010002.
- Scherer, M., Otto, V., Stump, J.D., Klingl, S., Müller, R., Reuter, N., Muller, Y.A., Sticht, H., Stamminger, T. (2015). Characterization of recombinant human cytomegaloviruses encoding IE1 mutants L174P and 1-382 reveals that viral targeting of PML bodies perturbs both intrinsic and innate immune responses. J. Virol. 90(3): 1190-205.
- Thomas, M., Sonntag, E., Müller, R., Schmidt, S., Zielke, B., Fossen, T., Stamminger, T. (2015). pUL69 of human cytomegalovirus recruits the cellular protein arginine methyltransferase 6 via a domain that is crucial for mRNA export and efficient viral replication. J. Virol. 89(18): 9601-15.
- Klingl, S., Scherer, M., Stamminger, T., Muller, Y.A. (2015). Controlled crystal dehydration triggers a space-group switch and shapes the tertiary structure of cytomegalovirus immediate-early 1 (IE1) protein. Acta Crystallogr. D Biol Crystallogr. 71: 1493-504.
- Wagenknecht, N., Reuter, N., Scherer, M., Reichel, A., Müller, R., Stamminger, T. (2015). Contribution oft he major ND10 proteins PML, hDaxx and Sp100 to the regulation of human cytomegalovirus latency and lytic replication in the monocytic cell line THP-1. Viruses 7(6): 2884-907.
- Vink, El, Zheng, Y., Yeasmin, R., Stamminger, T., Krug, L.T., Hearing, P. (2015). Impact of adenovirus E4-ORF3 oligomerization and protein localization on cellular gene expression. Viruses 7(5): 2428-49.
- Scherer, M., Klingl, S., Sevvana, M., Otto, V., Schilling, E.M., Stump, J.D., Müller, R., Reuter, N., Sticht, H., Muller, Y.A., Stamminger, T. (2014). Crystal structure of cytomegalovirus IE1 protein reveals targeting of TRIM family member PML via coiled-coil interactions. PLoS Pathog. 10(11): e1004512.
Reviews
- Scherer, M., Schilling, E.M., Stamminger, T. (2017) The human CMV IE1 protein: an offender of PML nuclear bodies. Adv. Anat. Embryol. Cell Biol., 223: 77-94.
- Scherer, M., and Stamminger, T. (2016). Emerging role of PML nuclear bodies in innate immune signaling. J. Virol., 90(13): 5840-4.
- Scherer, M., Wagenknecht, N., Reuter, N., Stamminger, T.(2016). Silencing of human cytomegalovirus gene expression mediated by components of PML nuclear bodies. In: Epigenetics – a different way of looking at genetics (Ed.: W. Dörfler, P. Böhm). Springer, Heidelberg, Germany, pp 175-196.
- Scherer, M., Stamminger, T. (2014). The human cytomegalovirus IE1 protein – past and present developments. Future Virology. 9(4): 415-430.
- Thomas, M., Reuter, N., Stamminger, T. (2013). Multifacted regulation of human cytomegalovirus gene expression. In: Cytomegaloviruses – from molecular biology to intervention (Ed.: M. Reddehase). Caister Academic Press, Norfolk, UK.
Research group of Prof. Dr. Jens von Einem
Regulation of HCMV morphogenesis
Our research focuses on the cytoplasmic steps of human cytomegalovirus (HCMV) morphogenesis and their regulation by viral and cellular proteins. The generation and release of infectious virus particles require the acquisition of the viral envelope, a cell derived lipid bilayer with embedded viral glycoproteins. This envelope is obtained as a result of a cytoplasmic budding process of tegumented capsids into vesicles containing viral glycoprotein complexes, known as secondary envelopment. Because of its essential role for HCMV infection and the need of novel antiviral drugs, we aim to identify and characterize the molecular mechanisms driving secondary envelopment and explore potential targets that could be used to limit HCMV infection.

Secondary envelopment: a well-coordinated multistep process. HCMV secondary envelopment takes place in a particular cytoplasmic region, termed the viral assembly complex (vAC). The formation of the vAC in cells is induced by infection and involves a remarkable reorganization of various cellular membranes including those of the endoplasmic reticulum, the Golgi intermediate compartment, the Golgi compartment and the endosomal pathways.
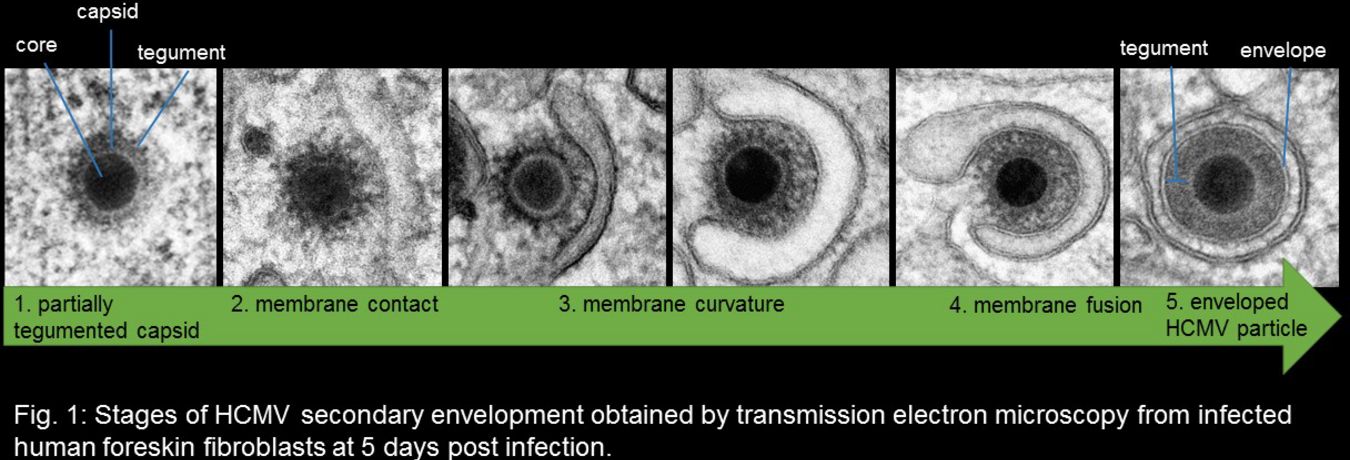
HCMV can productively infect various different cell types, including endothelial cells, epithelial cells, fibroblasts and macrophages. However, vAC formation and virus morphogenesis stages are similar in these different cell types. Notably, the vAC is the only cytoplasmic site in HCMV infected cells where secondary envelopment occurs. Accordingly, many viral components (glycoproteins, tegument proteins and capsids), known to be involved in viral assembly, localize and accumulate at the vAC during infection. The ideal method for visualization of the virus morphogenesis and especially of secondary envelopment is electron microscopy (EM). Therefore, we work in close collaboration with the Central Facility for Electron Microscopy (Ulm University). Novel fixation methods in combination with advanced electron microscopic techniques (e.g. 3D-electron microscopy) are used to analyse HCMV infected cells. These methods are specifically well adapted for the preservation and study of membrane structures and enables us to quantify secondary envelopment stages in infected cells.
Secondary envelopment can be divided into several steps at which different subsets of viral or cellular proteins seem to be involved (Fig. 1). To initiate the envelopment process, partially tegumented capsids need to be directed to the vAC where they establish contact to cellular membranes that contain the viral glycoprotein complexes. We could recently identify HCMV tegument protein pUL47 to function in targeting of partially tegumented capsids to the vAC. This protein forms a complex with the large tegument protein pUL48 and thus may act in complex to target capsids to the vAC and to membranes where secondary envelopment takes place. These membranes subsequently wrap around the capsid. This process involves the induction of membrane curvature. The tegument layer, as found in mature virions, starts to form concomitant with progressing envelopment. These processes require the presence of the pUL99/pUL94 tegument protein complex. We are currently investigating the mechanism of intracellular trafficking of the pUL99/pUL94 complex and the contribution of individual complex partners to the function of this complex. Ultimately, a membrane fusion process occurs, resulting in the viral envelope and an additional membrane enclosing the virus particle.
Together, our EM studies of HCMV morphogenesis propose secondary envelopment as a highly coordinated process at the vAC that is regulated by several viral proteins. For most of these proteins, functional homologs can be found in other herpesviruses, suggesting conserved domains and mechanisms. Targeting of these mechanisms could therefore provide measures against several herpesviruses.
Regulation of membrane fusion. Secondary envelopment is completed by a membrane fusion event by which the viral envelope is generated. Recently, we have made achievements towards a more detailed understanding of this membrane fusion process. We have identified that the membrane fusion process requires HCMV tegument protein pUL71. The absence of pUL71 resulted in an increased number of capsids at the vAC that were in contact with or partially engulfed (surrounded) by membranes. It thus appears that directing capsids to membranes and initiation of the envelopment process was still possible whereas the final steps of envelopment, involving membrane fusion, were severely reduced. We could further show in a collaborative project with the lab of Elke Bogner (Charité Berlin), that pUL71 forms oligomers via an N-terminal leucine-zipper motif and that the leucine-zipper is required for the function of pUL71 during secondary envelopment. It is also known that pUL71 forms a complex with tegument protein pUL103. Current research concentrates on clarification of the exact role of pUL103 in membrane fusion, to investigate the role of the pUL71/pUL103 complex in detail and to elucidate the mechanism of HCMV membrane fusion.
Intracellular trafficking of viral proteins. The challenge for herpesviruses is the temporal and spatial coordination of the many viral structural proteins that need to be incorporated into virions. Different intracellular trafficking mechanisms were already shown to be used by various HCMV encoded proteins for their accumulation at the vAC. We could recently show that trafficking of pUL71 to the vAC is regulated by utilizing host trafficking mechanisms via the presence of a N-terminal tyrosine-based trafficking motif (YxxΦ) in pUL71. Endocytosis of HCMV pUL71 via the YxxΦ motif may allow the retrieval of this protein for the process of secondary envelopment. Similar trafficking motifs could also be found in cytoplasmic domains of several viral glycoproteins. The utilization of the same intracellular trafficking pathways by several viral proteins may represent a possible mechanism to concentrate these proteins at the site of secondary envelopment. Thus, intracellular trafficking, if targeted, might represent a novel target for antiviral therapy.
Selected References
Dietz AN, Villinger C, Becker S, Frick M, von Einem J (2018) A Tyrosine-Based Trafficking Motif of the Tegument Protein pUL71 Is Crucial for Human Cytomegalovirus Secondary Envelopment. J Virol 92:e00907-17 . doi: 10.1128/JVI.00907-17
Biolatti M, Dell’Oste V, Pautasso S, Gugliesi F, von Einem J, Krapp C, Jakobsen MR, Borgogna C, Gariglio M, De Andrea M, Landolfo S (2017) The Human Cytomegalovirus Tegument Protein pp65 (pUL83) Dampens Type I Interferon Production by Inactivating the DNA Sensor cGAS without Affecting STING. J Virol JVI.01774-17 . doi: 10.1128/JVI.01774-17
Biolatti M, Dell’Oste V, Pautasso S, von Einem J, Marschall M, Plachter B, Gariglio M, De Andrea M, Landolfo S (2016) Regulatory Interaction between the Cellular Restriction Factor IFI16 and Viral pp65 (pUL83) Modulates Viral Gene Expression and IFI16 Protein Stability. J Virol 90:JVI.00923-16 . doi: 10.1128/JVI.00923-16
Lump E, Castellano LM, Meier C, Seeliger J, Erwin N, Sperlich B, Stürzel CM, Usmani S, Hammond RM, von Einem J, Gerold G, Kreppel F, Bravo-Rodriguez K, Pietschmann T, Holmes VM, Palesch D, Zirafi O, Weissman D, Sowislok A, Wettig B, Heid C, Kirchhoff F, Weil T, Klärner F-G, Schrader T, Bitan G, Sanchez-Garcia E, Winter R, Shorter J, Münch J (2015) A molecular tweezer antagonizes seminal amyloids and HIV infection. Elife 4:e05397 . doi: 10.7554/eLife.05397
Cappadona I, Villinger C, Schutzius G, Mertens T, von Einem J (2015) Human Cytomegalovirus pUL47 Modulates Tegumentation and Capsid Accumulation at the Viral Assembly Complex. J Virol 89:7314–28 . doi: 10.1128/JVI.00603-15
Bogdanow B, Weisbach H, von Einem J, Straschewski S, Voigt S, Winkler M, Hagemeier C, Wiebusch L (2013) Human cytomegalovirus tegument protein pp150 acts as a cyclin A2-CDK-dependent sensor of the host cell cycle and differentiation state. Proc Natl Acad Sci U S A 110:17510–5 . doi: 10.1073/pnas.1312235110
Wang D, Li G, Schauflinger M, Nguyen CC, Hall ED, Yurochko AD, von Einem J, Kamil JP (2013) The ULb’ region of the human cytomegalovirus genome confers an increased requirement for the viral protein kinase UL97. J Virol 87:6359–76 . doi: 10.1128/JVI.03477-12
Brock I, Krüger M, Mertens T, von Einem J (2013) Nuclear targeting of human cytomegalovirus large tegument protein pUL48 is essential for viral growth. J Virol 87:6005–19 . doi: 10.1128/JVI.03558-12
Schauflinger M, Villinger C, Mertens T, Walther P, von Einem J (2013) Analysis of human cytomegalovirus secondary envelopment by advanced electron microscopy. Cell Microbiol 15:305–314 . doi: 10.1111/cmi.12077
Meissner CS, Suffner S, Schauflinger M, von Einem J, Bogner E (2012) A leucine zipper motif of a tegument protein triggers final envelopment of human cytomegalovirus. J Virol 86:3370–82 . doi: 10.1128/JVI.06556-11
Maninger S, Bosse JB, Lemnitzer F, Pogoda M, Mohr CA, von Einem J, Walther P, Koszinowski UH, Ruzsics Z (2011) M94 is essential for the secondary envelopment of murine cytomegalovirus. J Virol 85:9254–67 . doi: 10.1128/JVI.00443-11
Schauflinger M, Fischer D, Schreiber A, Chevillotte M, Walther P, Mertens T, von Einem J (2011) The tegument protein UL71 of human cytomegalovirus is involved in late envelopment and affects multivesicular bodies. J Virol 85:3821–32 . doi: 10.1128/JVI.01540-10
Chevillotte M, von Einem J, Meier BM, Lin F-M, Kestler HA, Mertens T (2010) A new tool linking human cytomegalovirus drug resistance mutations to resistance phenotypes. Antiviral Res 85:318–327 . doi: 10.1016/J.ANTIVIRAL.2009.10.004
Chevillotte M, Schubert A, Mertens T, von Einem J (2009) Fluorescence-based assay for phenotypic characterization of human cytomegalovirus polymerase mutations regarding drug susceptibility and viral replicative fitness. Antimicrob Agents Chemother 53:3752–61 . doi: 10.1128/AAC.00165-09
Chevillotte M, Landwehr S, Linta L, Frascaroli G, Lüske A, Buser C, Mertens T, von Einem J (2009) Major tegument protein pp65 of human cytomegalovirus is required for the incorporation of pUL69 and pUL97 into the virus particle and for viral growth in macrophages. J Virol 83:2480–90 . doi: 10.1128/JVI.01818-08
Tischer BK, Von Einem J, Kaufer B, Osterrieder N (2006) Two-step Red-mediated recombination for versatile high-efficiency markerless DNA manipulation in Escherichia coli. Biotechniques 40:191–197 . doi: 10.2144/000112096
Funding
Boehringer Ingelheim Ulm University (BIU) BioCenter
Junior Professor Program of the MWK Baden-Wuerttemberg, “Viral morphogenesis as target for antiviral intervention ….”
Bausteinförderung 3.2 der Medizinischen Fakultät (Dr. Clarissa Villinger)
Research group of Prof. Dr. Christian Sinzger
Entry of HCMV into target cells: underlying mechanisms and development of inhibitors
Human cytomegalovirus HCMV (HCMV) belongs to the family of herpesviruses. Usually, it is acquired early in life via breastfeeding or contact among toddlers and then persists in the organism for a lifetime. In immunocompetent hosts, the infection often goes unnoticed or causes a transient mild disease. In contrast, mother to fetus transmission during a pregnancy can damage the brain and sensory organs, sensorineural hearing loss being the most frequent manifestation. For AIDS patients, an HCMV infection is hazardous as it may cause retinitis and finally blindness. In transplant recipient who are immunosuppressed due to their treatment regimen, HCMV may affect the lung, the gastrointestinal tract or the transplanted organ.
We are investigating, which cells are infected by HCMV in the organism, how the virus enters into these cells and how this can be prevented. Neutralization of virus by antibodies is a common approach to block virus entry, and we try to further optimize this strategy for HCMV. In addition, we are trying to resolve the molecular virus-cell interactions contributing to HCMV entry as a basis for the development of novel direct-acting antiviral agents that interfere with adsorption and/or penetration of this virus.
Main projects:
- Identification of highly and broadly neutralizing antibodies against HCMV
- Role of viral glycoprotein complexes for entry of HCMV into various cell types
- Derivatives of cellular surface proteins for inhibition of HCMV entry
Lab members:
Christian Sinzger, Prof. Dr. med. (Principal Investigator); Cora Stegmann, PhD student; Jessica Julia Falk, PhD student; Kerstin Laib Sampaio, technician; Dagmar Stöhr, technician; Dina Fischer, MD student; Chiara Antonini, master student
Highly and broadly neutralizing antibodies against HCMV. Following an infection with HCMV, our immune system produces antibodies against this virus. However, this antibody response does not reliably prevent subsequent reactivation of the virus, mother-to-child transmission, and the development of disease under conditions of reduced cellular immunity. We hypothesize that this partial failure is due to (i) only low neutralizing activity against infection of connective tissue cells and (ii) lack of activity against cell-to-cell spread of the virus. Therefore we screen blood donor populations in order to identify “elite neutralizers” who produce exceptionally potent antibodies that might provide a basis for improved antibody-based treatment of HCMV infections.
To achieve this, we developed a two-step screening approach and tested plasma samples of 10,000 HCMV-seropositive blood donors for high neutralizing activity against a Gaussia-luciferase expressing reporter virus. Top neutralizers were further tested by an ELISA-based neutralization assay regarding broad efficiency against different HCMV strains. Donors with highly and broadly neutralizing plasma are regarded “elite neutralizers”. We are currently investigating (i) whether plasma from such donors can be directly used to improve the treatment of transplant recipients and (ii) whether they can serve as a source of therapeutic monoclonal antibodies. This project is a collaboration with Prof. Ramin Lotfi (Transfusion Medicine, Ulm, Germany) and Dr. Ady Krawczyk (Virology, Essen Germany).
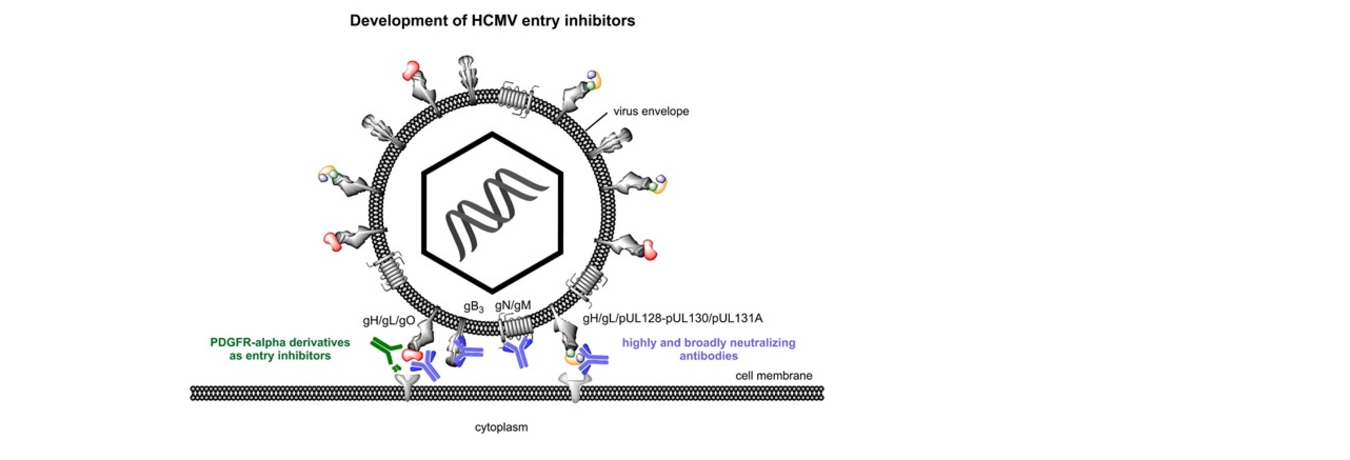
Fig. 1: Strategies for the development of entry inhibitors for the treatment of HCMV infections
Role of viral glycoprotein complexes for entry of HCMV. The capacity of HCMV to infect a variety of different cell types is regarded an important factor for the pathogenicity of this virus. The presence of certain glycoprotein complexes in the virus envelope have been identified as the molecular basis of this broad cell tropism. In our lab, we use fibroblasts and endothelial cells as model cell types representing two different entry pathways
the trimeric complex gH/gL/gO is necessary for efficient entry into both cell types
in addition, the pentameric complex gH/gL/pUL128/pUL130/pUL131A is necessary for infection of endothelial cells
Both envelope protein complexes contain the essential viral proteins gH and gL, which are assumed to contribute to virus penetration by triggering the fusogenic activity of another viral envelope protein complex, the gB homotrimer. Interestingly, the ratio of trimer and pentamer in the virus envelope greatly varies between different laboratory strains of HCMV, which may influence infection efficiency and cell tropism of the respective viruses. Using mutational approaches we attempt to decipher the exact contribution of the two alternative gH/gL complexes to HCMV entry and spread at the molecular level.
Derivatives of cellular surface proteins for inhibition of HCMV entry. We and others have found that the trimeric viral envelope protein complex gH/gL/gO binds to the cellular surface protein PDGFR-alpha (platelet derived growth factor receptor alpha), and that this interaction is necessary for efficient entry. Based on this finding, we have identified a soluble form of this cellular HCMV receptor, PDGFR-alpha-Fc, as a potent inhibitor of HCMV entry into both model cell types, fibroblasts and endothelial cells. We are currently investigating the therapeutic potential of such PDGFR-alpha derivatives.
In addition, we have identified three PDGFR-alpha derived 40mer peptides that also inhibit HCMV entry. We are now investigating the exact mode of action of these peptides and try to further optimize their efficacy by targeted modifications. We are particularly interested in developing inhibitory PDGFR-alpha-derived peptides that can block both cell free entry and cell-to-cell spread of HCMV in a variety of cell types.
Collaborations
Prof. Dr. Ramin Lotfi, Institute for Transfusion Medicine, University Hospital Ulm, Germany
Dr. Adalbert Krawczyk, Institute of Virology, University Hospital Essen, Germany
Prof. Dr. Paul Walter, Central Facility for Electron Microscopy, Ulm University, Germany
Prof. Dr. Michael Mach Institute of Clinical and Molecular Virology, University of Erlangen-Nürnberg, Germany
Prof. Dr. Bodo Plachter, Institute for Virology, University Medical Center of the Johannes Gutenberg University Mainz, Germany
PD Dr. Barbara Adler, Max von Pettenkofer-Institute for Virology, Ludwig-Maximilians-University Munich, Germany
Dr. Peter Lischka, AiCuris GmbH & Co KG, Wuppertal, Germany
Prof. Richard J. Stanton, Institute of Infection and Immunity, School of Medicine, Cardiff University, UK
Prof. Dr. Renata Stripecke, Clinics of Hematology, Hemostasis, Oncology and Stem Cell Transplantation, Hannover Medical School, Hannover, Germany
Funding
Else Kröner-Fresenius-Stiftung, Projekt Nr. 2016-A126
Selected References
Stegmann C, Hochdorfer D, Lieber D, Subramanian N, Stöhr D, Laib Sampaio K, Sinzger C. A derivative of platelet-derived growth factor receptor alpha binds to the trimer of human cytomegalovirus and inhibits entry into fibroblasts and endothelial cells. Plos Pathogens. 2017, 13, e1006273.
Sampaio KL, Weyell A, Subramanian N, Wu Z, Sinzger C. A TB40/E-derived human cytomegalovirus genome with an intact US-gene region and a self-excisable BAC cassette for immunological research. Biotechniques. 2017, 63: 205-214. doi: 10.2144/000114606. PubMed PMID: 29185920
Falk JJ, Winkelmann M, Schrezenmeier H, Sinzger C, Lotfi R. A two-step screening approach for the identification of blood donors with highly and broadly neutralizing capacities against human cytomegalovirus. submitted Transfusion. 2017, 57: 412-422..
Maier AK, Jung R, Villinger C, Schubert A, Walther P, Sinzger C, Lieber D. A luciferase gene driven by an alphaherpesviral promoter also responds to immediate early antigens of the betaherpesvirus hcmv, allowing comparative analyses of different human herpesviruses in one reporter cell line. PLoS One. 2017, 12:e0169580.
Stegmann, C, Abdellatif MEA, Laib Sampaio, K, Walther P, Sinzger, C. Importance of highly conserved peptide sites of HCMV gO for the formation of the gH/gL/gO complex. J Virol. 2017, 91: e01339-16.
Falk JJ, Laib Sampaio K, Stegmann C, Lieber D, Kropff B, Mach M, Sinzger C. Generation of a Gaussia luciferase-expressing endotheliotropic cytomegalovirus for screening approaches and mutant analyses. J Virol Methods. 2016, 17: 235:182-9.
Hochdorfer D, Florin L, Sinzger C, Lieber D. Tetraspanin CD151 Promotes Initial Events in Human Cytomegalovirus Infection. J Virol. 2016, 90: 6430-42.
Laib Sampaio K, Stegmann C, Brizic I, Adler B, Stanton RJ, Sinzger C. The contribution of pUL74 to growth of human cytomegalovirus is masked in the presence of RL13 and UL128 expression. J Gen Virol. 2016. 97: 1917-27.
Lieber D, Hochdorfer D, Stoehr D, Schubert A, Lotfi R, May T, Wirth D, Sinzger C. A permanently growing human endothelial cell line supports productive infection with human cytomegalovirus under conditional cell growth arrest. BioTechniques. 2015, 59:127-36.
Sampaio KL, Jahn G, Sinzger C. Applications for a dual fluorescent human cytomegalovirus in the analysis of viral entry. Methods Mol Biol. 2013;1064:201-9.
Schuessler, A., Laib Sampaio, K. Sinzger, C. Charge cluster-to-alanine scanning of UL128 for fine tuning of the endothelial cell tropism of human cytomegalovirus. J Virol, 2008, 82, 11239-11246.
Sinzger, C., Hahn, G., Digel, M., Katona, R., Laib Sampaio, K., Messerle, M., Hengel, H., Koszinowski, U., Brune, W. and Adler, B. (2008). Cloning and sequencing of a highly productive, endotheliotropic virus strain derived from human cytomegalovirus TB40/E. J Gen Virol. 89: 359-368.
Laib Sampaio, K., Cavignac, Y., Stierhof, Y. D. and Sinzger, C. Human cytomegalovirus labeled with green fluorescent protein for live analysis of intracellular particle movements. J Virol. 2005, 79: 2754-2767
Riegler, S., Hebart, H., Einsele, H., Brossart, P., Jahn, G. and Sinzger, C. Monocyte-derived dendritic cells are permissive to the complete replicative cycle of human cytomegalovirus. J Gen Virol. 2000, 81: 393-399
Sinzger, C., Schmidt, K., Knapp, J., Kahl, M., Beck, R., Waldman, J., Hebart, H., Einsele, H. and Jahn, G. Modification of human cytomegalovirus tropism through propagation in vitro is associated with changes in the viral genome. J Gen Virol. 1999, 80: 2867-2877.
Sinzger, C., Plachter, B., Grefte, A., The, T. H. and Jahn, G. Tissue macrophages are infected by human cytomegalovirus in vivo. J Infect Dis. 1996, 173: 240-245
Sinzger, C., Grefte, A., Plachter, B., Gouw, A. S., The, T. H. and Jahn, G. (1995). Fibroblasts, epithelial cells, endothelial cells and smooth muscle cells are major targets of human cytomegalovirus infection in lung and gastrointestinal tissues. J Gen Virol. 1995, 76: 741-750
Reviews
- Frascaroli G, Sinzger C. Distinct properties of human cytomegalovirus strains and the appropriate choice of strains for particular studies. Methods in molecular biology. 2014;1119:29-46.
- Adler B, and Sinzger C. Cytomegalovirus Inter-Strain Variance in Cell-Type Tropism. In Cytomegaloviruses: From Molecular Pathogenesis to Intervention. M. J. Reddehase (ed.), 2013 Caister Academic Press, vol. 1 , 297-321.
Research group of Prof. Dr. Detlef Michel
The main topics of our research are the improvement, development or establishment of new diagnostic methods and the characterization of human cytomegalovirus drug resistance mechanisms. Consequently, we are conducting and optimizing phenotypic and genotypic assays to evaluate antiviral resistance of different viruses (e.g. HSV-1, HCMV, HBV, HIV) against a variety of antiviral drugs. Furthermore, we offer and constantly refine a bioinformatics tool, termed mutation resistance analyser (MRA) →, that allows the prediction of drug-resistance for HCMV, HSV and HCV
Selected References
- Müller JA., Harms M., Schubert A., Mayer B., Jansen S., Herbeuval JP., Michel D., Mertens T., Vapalahti O., Schmidt-Chanasit J., and Münch J.(2017) Development of a high-throughput colorimetric Zika virus infection assay. Med Microbiol Immunol. 206 (2):175-185.
- Böttcher S, Obermeier PE, Neubauer K, Diedrich S; the Laboratory Network for Enterovirus Diagnostics. (2016) Recombinant enterovirus A71 subgenogroup C1 strains, Germany, 2015. Emerg Infect Dis. 22 (10), DOI: 10.3201
- Müller J.A., Harms M., Schubert A., Jansen S., Michel D., Mertens T., Schmidt-Chanasit J, and Münch J. (2016) Inactivation and Environmental Stability of Zika Virus. Emerg Infect Dis. 22 (9). doi: 10.3201
- Lischka P., Michel D., and Zimmermann H. (2016) Characterization of Cytomegalovirus Breakthrough Events in a Phase 2 Prophylaxis Trial of Letermovir (AIC246, MK 8228). J. Infect. Dis. 213 (1): 23-30
- Michel D. (2014) Influenzaverdacht, ist der Labortest Pflicht? Der Allgemeinarzt 36 (10): 16-21.
- de Paula V.S., Perse da Silva A., Michel D., Melgaço J.G., Vieira Y.R., Dos Santos D.C., Carvalho Viana de Araújo C., Pacheco-Moreira L.F., Pelajo-Machado M., do Nascimento C.M., Schmidt-Chanasit J., Alves Pinto M. (2014) Acute liver failure in an immunocompetent patient. J Clin Virol. 22. pii: S1386-6532 (14) 10.1016
- Stoelben S., Arns W., Renders L., Hummel J., Mühlfeld A., Stangl M., Fischereder M., Gwinner W., Suwelack B., Witzke O., Dürr M., Beelen D., Michel D., Lischka P., Zimmermann H., Rübsamen-Schaeff H., Budde K. (2014) Pre-emptive treatment of Cytomegalovirus infection in kidney transplant recipients with letermovir: results of a Phase 2a study. Transpl Int. 27 (1):77-86.
- Bommer M. and Michel D. (2013) Prevention of cytomegalovirus disease in patients with impaired cell mediated immunity – is there a need for maribavir? Expert Opinion on Orphan Drugs 1 (10): 829–836.
- Schubert A., Ehlert K., Schuler-Luettmann S., Gentner E., Mertens T., Michel D. (2013) Fast selection of maribavir resistant cytomegalovirus in a bone marrow transplant recipient. BMC Infect Dis. 13 (1):330.
- Schubert A., Michel D., Mertens T. (2013)Late HBsAg seroreversion of mutated hepatitis B virus after bone marrow transplantation. BMC Infect Dis. 13 (1):223.
- Wagner S., Arnold F., Wu Z., Schubert A., Walliser C., Tadagaki K., Jockers R., Mertens T., Michel D. (2012) The 7-transmembrane protein homologue UL78 of the human cytomegalovirus forms oligomers and traffics between the plasma membrane and different intracellular compartments. Arch Virol. 157 (5):935-949.
- Härter G., Michel D. (2012) Antiviral treatment of cytomegalovirus infection: an update. Expert Opin Pharmacother. 13 (5):623-627.
- Frenzel K., Ganepola S., Michel D., Thiel E., Krüger D.H., Uharek L., Hofmann J. (2012) Antiviral function and efficacy of polyvalent immunoglobulin products against CMV isolates in different human cell lines. Med Microbiol Immunol. 201 (3):277-286.
2018 | Chromatin-Remodeling Factor SPOC1 Acts as a Cellular Restriction Factor against Human Cytomegalovirus by Repressing the Major Immediate Early Promoter. | Reichel A, Stilp AC, Scherer M, Reuter N, Lukassen S, Kasmapour B, Schreiner S, Cicin-Sain L, Winterpacht A, Stamminger T |
2018 | Signatures of T and B Cell Development, Functional Responses and PD-1 Upregulation After HCMV Latent Infections and Reactivations in Nod. Rag. Gamma Mice Humanized With Cord Blood CD34(+) Cells | Front Immunol 2018; 9() |
2018 | Detection of antibody-secreting cells specific for the cytomegalovirus and herpes simplex virus surface antigens. | J Immunol Methods 2018; 462(): 13-22 |
2018 | A Tyrosine-Based Trafficking Motif of the Tegument Protein pUL71 Is Crucial for Human Cytomegalovirus Secondary Envelopment. | J Virol 2018; 92(1) |
2018 | Semen inhibits Zika virus infection of cells and tissues from the anogenital region. | Nat Commun 2018; 9(1): 2207 |
2018 | The molecular tweezer CLR01 inhibits Ebola and Zika virus infection. | Antiviral Res 2018; 152(): 26-35 |
2018 | Identification of Elite Neutralizers With Broad and Potent Neutralizing Activity Against Human Cytomegalovirus (HCMV) in a Population of HCMV-Seropositive Blood Donors. | J Infect Dis 2018; 218(6): 876-885 |
2018 | Large-Scale Screening of HCMV-Seropositive Blood Donors Indicates that HCMV Effectively Escapes from Antibodies by Cell-Associated Spread. | Viruses 2018; 10(9) |
2018 | Human Cytomegalovirus Tegument Protein pp65 (pUL83) Dampens Type I Interferon Production by Inactivating the DNA Sensor cGAS without Affecting STING. | J Virol 2018; 92(6) |
2018 | Formation and mosaicity of coccolith segment calcite of the marine algae Emiliania huxleyi. | J Phycol 2018; 54(1): 85-104 |
2018 | Investigating HCMV entry into host cells by STEM tomography. | J Struct Biol 2018; 204(3): 406-419 |
2018 | Inhibition of Tetraspanin Functions Impairs Human Papillomavirus and Cytomegalovirus Infections. | Int J Mol Sci 2018; 19(10) |
2018 | Classic paper: Are the chickenpox virus and the zoster virus identical?: HELMUT RUSKA. | Rev Med Virol 2018; 28(3): e1975 |
2018 | A Virally Encoded DeSUMOylase Activity Is Required for Cytomegalovirus Reactivation from Latency. | Cell Rep 2018; 24(3): 594-606 |
2018 | Human Macrophages Escape Inhibition of Major Histocompatibility Complex-Dependent Antigen Presentation by Cytomegalovirus and Drive Proliferation and Activation of Memory CD4 and CD8 T Cells. | Front Immunol 2018; 9(): 1129 |
2018 | Varicella-zoster-virus vaccination in immunosuppressed children with rheumatic diseases using a pre-vaccination check list | Pediatr Rheumatol Online J 2018; 16(1): 15 |
2017 | Tetraspanins in infections by human cytomegalo- and papillomaviruses. | Biochem Soc Trans. 2017;45(2): 489-497 (Impact(2016)=2.765, Typ=Review; Journal Article; Review; Research Support, Non-U.S. Gov't) |
2017 | A TB40/E-derived human cytomegalovirus genome with an intact US-gene region and a self-excisable BAC cassette for immunological research. | Biotechniques. 2017;63(5): 205-214 (Impact(2016)=2.03, Typ=Article; Journal Article) |
2017 | Naïve T cells are activated by autologous HCMV-infected endothelial cells through NKG2D and can control HCMV transmission in vitro. | J Gen Virol. 2017;98: 3068-3085 (Impact(2016)=2.838, Typ=Journal Article) |
2017 | Importance of Highly Conserved Peptide Sites of Human Cytomegalovirus gO for Formation of the gH/gL/gO Complex. | J Virol. 2017;91(1) (Impact(2016)=4.663, Typ=Journal Article) |
2017 | Human Cytomegalovirus Particles Treated with Specific Antibodies Induce Intrinsic and Adaptive but Not Innate Immune Responses. | J Virol. 2017;91(22) (Impact(2016)=4.663, Typ=Journal Article) |
2017 | Development of a high-throughput colorimetric Zika virus infection assay. | Med Microbiol Immunol (Berl). 2017;206(2): 175-185 (Impact(2016)=3.093, Typ=Journal Article; Journal Article) |
2017 | A Luciferase Gene Driven by an Alphaherpesviral Promoter Also Responds to Immediate Early Antigens of the Betaherpesvirus HCMV, Allowing Comparative Analyses of Different Human Herpesviruses in One Reporter Cell Line. | PLoS ONE. 2017;12(1): e0169580 (Impact(2016)=2.806, Typ=Article; Journal Article) |
2017 | A derivative of platelet-derived growth factor receptor alpha binds to the trimer of human cytomegalovirus and inhibits entry into fibroblasts and endothelial cells. | PLoS Pathog. 2017;13(4): e1006273 (Impact(2016)=6.608, Typ=Journal Article) |
2017 | Peptide vaccination in the presence of adjuvants in patients after hematopoietic stem cell transplantation with CD4+ T cell reconstitution elicits consistent CD8+ T cell responses. | Theranostics. 2017;7(6): 1705-1718 (Impact(2016)=8.712, Typ=Article; Journal Article) |
2017 | A two-step screening approach for the identification of blood donors with highly and broadly neutralizing capacities against human cytomegalovirus. | Transfusion. 2017;57(2): 412-422 (Impact(2016)=3.386, Typ=Journal Article) |
2017 | Antibody-based immunotherapy of aciclovir resistant ocular herpes simplex virus infections. | Virology. 2017;512: 194-200 (Impact(2016)=3.353, Typ=Article; Journal Article; Research Support, Non-U.S. Gov't) |
2016 | Two Cases of Vaccine-Derived Poliovirus Infection in an Oncology Ward. | N Engl J Med. 2016;374(13): 1296-8 (Impact(2015)=59.558, Typ=Letter; Letter; Case Reports) |
2016 | Generation of a Gaussia luciferase-expressing endotheliotropic cytomegalovirus for screening approaches and mutant analyses. | J Virol Methods. 2016;235: 182-9 (Impact(2015)=1.508, Typ=Article; Journal Article) |
2016 | Tetraspanin CD151 Promotes Initial Events in Human Cytomegalovirus Infection. | J Virol. 2016;90(14): 6430-42 (Impact(2015)=4.606, Typ=Article; Journal Article) |
2016 | Regulatory Interaction between the Cellular Restriction Factor IFI16 and Viral pp65 (pUL83) Modulates Viral Gene Expression and IFI16 Protein Stability. | J Virol. 2016;90(18): 8238-50 (Impact(2015)=4.606, Typ=Article; Journal Article) |
2016 | Characterization of Cytomegalovirus Breakthrough Events in a Phase 2 Prophylaxis Trial of Letermovir (AIC246, MK 8228). | J Infect Dis. 2016;213(1): 23-30 (Impact(2015)=6.344, Typ=Article; Journal Article) |
2016 | Phenotypic characterization of human cytomegalovirus strains in cell cultures based on their transmission kinetics. | J Gen Virol. 2016;97(9): 2376-86 (Impact(2015)=3.192, Typ=Article; Journal Article) |
2016 | The contribution of pUL74 to growth of human cytomegalovirus is masked in the presence of RL13 and UL128 expression. | J Gen Virol. 2016;97(8): 1917-27 (Impact(2015)=3.192, Typ=Journal Article) |
2016 | Megapinocytosis: a novel endocytic pathway. | Histochem Cell Biol. 2016;145(6): 617-27 (Impact(2015)=2.78, Typ=Article; Journal Article) |
2016 | Inactivation and Environmental Stability of Zika Virus. | Emerg Infect Dis. 2016;22(9): 1685-7 (Impact(2015)=6.994, Typ=Letter; Letter) |
2016 | Recombinant Enterovirus A71 Subgenogroup C1 Strains, Germany, 2015. | Emerg Infect Dis. 2016;22(10): 1843-6 (Impact(2015)=6.994, Typ=Letter; Letter) |
2016 | Identification of resistance-associated HCMV UL97- and UL54-mutations and a UL97-polymporphism with impact on phenotypic drug-resistance. | Antiviral Res. 2016;131: 1-8 (Impact(2015)=4.909, Typ=Article; Journal Article) |
2015 | A molecular tweezer antagonizes seminal amyloids and HIV infection. | eLife. 2015;4 (Impact(2015)=8.282, Typ=Journal Article; Research Support, Non-U.S. Gov't; Research Support, N.I.H., Extramural) |
2015 | 3D Analysis of HCMV Induced-Nuclear Membrane Structures by FIB/SEM Tomography: Insight into an Unprecedented Membrane Morphology. | Viruses. 2015;7(11): 5686-704 (Impact(2015)=3.042, Typ=Journal Article) |
2015 | The Cellular Proteins Grb2 and DDX3 Are Increased upon Human Cytomegalovirus Infection and Act in a Proviral Fashion. | PLoS ONE. 2015;10(6): e0131614 (Impact(2015)=3.057, Typ=Journal Article; Research Support, Non-U.S. Gov't) |
2015 | Natural killer cells can inhibit the transmission of human cytomegalovirus in cell culture by using mechanisms from innate and adaptive immune responses. | J Virol. 2015;89(5): 2906-17 (Impact(2015)=4.606, Typ=Journal Article) |
2015 | Interleukin-2 from Adaptive T Cells Enhances Natural Killer Cell Activity against Human Cytomegalovirus-Infected Macrophages. | J Virol. 2015;89(12): 6435-41 (Impact(2015)=4.606, Typ=Journal Article; Research Support, Non-U.S. Gov't) |
2015 | Human Cytomegalovirus pUL47 Modulates Tegumentation and Capsid Accumulation at the Viral Assembly Complex. | J Virol. 2015;89(14): 7314-28 (Impact(2015)=4.606, Typ=Journal Article; Research Support, Non-U.S. Gov't) |
2015 | A permanently growing human endothelial cell line supports productive infection with human cytomegalovirus under conditional cell growth arrest. | Biotechniques. 2015;59(3): 127-36 (Impact(2015)=2.298, Typ=Journal Article; Research Support, Non-U.S. Gov't) |
2014 | Schubert A, Gentner E, Bohn K, Schwarz M, Mertens T, Sauerbrei A. Antiviral Res. 2014 Jul;107:16-22. doi: 10.1016/j.antiviral.2014.03.015. Epub 2014 Apr 18. | |
2014 | Reinhardt B, Godfrey R, Fellbrich G, Frank H, Lüske A, Olieslagers S, Mertens T, Waltenberger J. Cardiovasc Res. 2014 Nov 1;104(2):315-25. doi: 10.1093/cvr/cvu204. Epub 2014 Sep 16. | |
2014 | Widespread context dependency of microRNA-mediated regulation. | Erhard F, Haas J, Lieber D, Malterer G, Jaskiewicz L, Zavolan M, Dölken L, Zimmer R. Genome Res. 2014 Jun;24(6):906-19. doi: 10.1101/gr.166702.113. Epub 2014 Mar 25. |
2014 | Flechtner-Mors M, Schick A, Oeztuerk S, Haenle MM, Wilhelm M, Koenig W, Imhof A, Boehm BO, Graeter T, Mason RA, Kratzer W, Akinli AS; EMIL-Study Group. Horm Metab Res. 2014 Apr;46(4):287-93. doi: 10.1055/s-0033-1354369. Epub 2013 Sep 2. | |
2014 | Paula VS, Silva AP, Michel D, Melgaço JG, Vieira YR, Santos DC, Araújo CC, Pacheco-Moreira LF, Pelajo-Machado M, Nascimento CM, Schmidt-Chanasit J, Pinto MA. J Clin Virol. 2014 May;60(1):1-3. doi: 10.1016/j.jcv.2014.03.008. Epub 2014 Mar 22. No abstract available. | |
2014 | Meisel R, Heseler K, Nau J, Schmidt SK, Leineweber M, Pudelko S, Wenning J, Zimmermann A, Hengel H, Sinzger C, Degistirici Ö, Sorg RV, Däubener W. Mediators Inflamm. 2014;2014:898630. doi: 10.1155/2014/898630. Epub 2014 Mar 23. | |
2014 | Frascaroli G, Sinzger C. Methods Mol Biol. 2014;1119:29-46. doi: 10.1007/978-1-62703-788-4_3. | |
2014 | Hübers A, Marroquin N, Schmoll B, Vielhaber S, Just M, Mayer B, Högel J, Dorst J, Mertens T, Just W, Aulitzky A, Wais V, Ludolph AC, Kubisch C, Weishaupt JH, Volk AE. Neurobiol Aging. 2014 May;35(5):1214.e1-6. doi: 10.1016/j.neurobiolaging.2013.11.034. Epub 2013 Dec 4. | |
2014 | Stoelben S, Arns W, Renders L, Hummel J, Mühlfeld A, Stangl M, Fischereder M, Gwinner W, Suwelack B, Witzke O, Dürr M, Beelen DW, Michel D, Lischka P, Zimmermann H, Rübsamen-Schaeff H, Budde K. Transpl Int. 2014 Jan;27(1):77-86. doi: 10.1111/tri.12225. Erratum in: Transpl Int. 2014 Apr;27(4):416. | |
2013 | Fischer L, Laib Sampaio K, Jahn G, Hamprecht K, Göhring K. Antiviral Res. 2013 Dec;100(3):575-7. doi: 10.1016/j.antiviral.2013.09.026. Epub 2013 Oct 10. | |
2013 | Late HBsAg seroreversion of mutated hepatitis B virus after bone marrow transplantation. | Schubert A, Michel D, Mertens T. BMC Infect Dis. 2013 May 16;13:223. doi: 10.1186/1471-2334-13-223. |
2013 | Fast selection of maribavir resistant cytomegalovirus in a bone marrow transplant recipient. | Schubert A, Ehlert K, Schuler-Luettmann S, Gentner E, Mertens T, Michel D. BMC Infect Dis. 2013 Jul 19;13:330. doi: 10.1186/1471-2334-13-330. |
2013 | Falkenhorst G, Harder T, Remschmidt C, Terhardt M, Zepp F, Ledig T, Wicker S, Keller-Stanislawski B, Mertens T. Bundesgesundheitsblatt Gesundheitsforschung Gesundheitsschutz. 2013 Nov;56(11):1557-64. doi: 10.1007/s00103-013-1844-9. Erratum in: Bundesgesundheitsblatt Gesundheitsforschung Gesundheitsschutz. 2014 Apr;57(4):473-4. | |
2013 | [Concepts, effectiveness, and perspectives of pandemic and seasonal influenza vaccines]. | Grund S, Wichmann O, Mertens T, Hengel H. Bundesgesundheitsblatt Gesundheitsforschung Gesundheitsschutz. 2013 Jan;56(1):76-86. doi: 10.1007/s00103-012-1590-4. German. |
2013 | Background paper to the recommendation for routine rotavirus vaccination of infants in Germany. | Koch J, Wiese-Posselt M, Remschmidt C, Wichmann O, Bertelsmann H, Garbe E, Hengel H, Meerpohl JJ, Mas Marques A, Oppermann H, Hummers-Pradier E, von Kries R, Mertens T. |
2013 | Analysis of human cytomegalovirus secondary envelopment by advanced electron microscopy. | Schauflinger M, Villinger C, Mertens T, Walther P, von Einem J. Cell Microbiol. 2013 Feb;15(2):305-14. doi: 10.1111/cmi.12077. Epub 2012 Dec 20. |
2013 | Mertens T. Dtsch Arztebl Int. 2013 Nov 22;110(47):791-2. doi: 10.3238/arztebl.2013.0791. No abstract available. | |
2013 | Bayer C, Varani S, Wang L, Walther P, Zhou S, Straschewski S, Bachem M, Söderberg-Naucler C, Mertens T, Frascaroli G. J Virol. 2013 Jan;87(1):67-79. doi: 10.1128/JVI.01585-12. Epub 2012 Oct 10. | |
2013 | Brock I, Krüger M, Mertens T, von Einem J. J Virol. 2013 May;87(10):6005-19. doi: 10.1128/JVI.03558-12. Epub 2013 Mar 20. | |
2013 | Wang D, Li G, Schauflinger M, Nguyen CC, Hall ED, Yurochko AD, von Einem J, Kamil JP. J Virol. 2013 Jun;87(11):6359-76. doi: 10.1128/JVI.03477-12. Epub 2013 Mar 27. | |
2013 | Wu Z, Sinzger C, Frascaroli G, Reichel J, Bayer C, Wang L, Schirmbeck R, Mertens T. J Virol. 2013 Jul;87(13):7717-25. doi: 10.1128/JVI.01096-13. Epub 2013 May 1. | |
2013 | Becker S, von Einem J. Methods Mol Biol. 2013;1064:29-41. doi: 10.1007/978-1-62703-601-6_3. | |
2013 | Determination of HSV-1 infectivity by plaque assay and a luciferase reporter cell line. | Lieber D, Bailer SM. Methods Mol Biol. 2013;1064:171-81. doi: 10.1007/978-1-62703-601-6_12. |
2013 | Generation of a stable cell line for constitutive miRNA expression. | Lieber D. Methods Mol Biol. 2013;1064:183-200. doi: 10.1007/978-1-62703-601-6_13. |
2013 | Applications for a dual fluorescent human cytomegalovirus in the analysis of viral entry. | Sampaio KL, Jahn G, Sinzger C. Methods Mol Biol. 2013;1064:201-9. doi: 10.1007/978-1-62703-601-6_14. |
2013 | Analysis of cell migration during human cytomegalovirus (HCMV) infection. | Varani S, Frascaroli G. Methods Mol Biol. 2013;1064:299-313. doi: 10.1007/978-1-62703-601-6_22. |
2013 | Assessment of natural killer cell responses to human cytomegalovirus-infected macrophages. | Wu Z, Frascaroli G, Mertens T. Methods Mol Biol. 2013;1064:289-98. doi: 10.1007/978-1-62703-601-6_21. |
2013 | Heseler K, Schmidt SK, Spekker K, Sinzger C, Sorg RV, Quambusch M, Zimmermann A, Meisel R, Däubener W. PLoS One. 2013 May 15;8(5):e64442. doi: 10.1371/journal.pone.0064442. Print 2013. | |
2013 | Neu C, Sedlag A, Bayer C, Förster S, Crauwels P, Niess JH, van Zandbergen G, Frascaroli G, Riedel CU. PLoS One. 2013 Jun 11;8(6):e66898. doi: 10.1371/journal.pone.0066898. Print 2013. | |
2013 | Bogdanow B, Weisbach H, von Einem J, Straschewski S, Voigt S, Winkler M, Hagemeier C, Wiebusch L. Proc Natl Acad Sci U S A. 2013 Oct 22;110(43):17510-5. doi: 10.1073/pnas.1312235110. Epub 2013 Oct 7. | |
2012 | Matysiak-Klose D, Ahmed F, Duclos P, Falck-Ytter Y, Forland F, Houweling H, Kramarz P, Langley JM, Mertens T, Schünemann H, Senouci K, Temte J, Wichmann O. Vaccine. 2012 Mar 23;30(14):2399-404. doi: 10.1016/j.vaccine.2011.12.004. Epub 2011 Dec 13. | |
2012 | NEX-TRAP, a novel method for in vivo analysis of nuclear export of proteins. | Raschbichler V, Lieber D, Bailer SM. Traffic. 2012 Oct;13(10):1326-34. doi: 10.1111/j.1600-0854.2012.01389.x. Epub 2012 Aug 1. |
2012 | A beta-herpesvirus with fluorescent capsids to study transport in living cells. | Bosse JB, Bauerfeind R, Popilka L, Marcinowski L, Taeglich M, Jung C, Striebinger H, von Einem J, Gaul U, Walther P, Koszinowski UH, Ruzsics Z. PLoS One. 2012;7(7):e40585. doi: 10.1371/journal.pone.0040585. Epub 2012 Jul 11. |
2012 | Frenzel K, Ganepola S, Michel D, Thiel E, Krüger DH, Uharek L, Hofmann J. Med Microbiol Immunol. 2012 Aug;201(3):277-86. doi: 10.1007/s00430-012-0229-2. | |
2012 | Schuessler A, Sampaio KL, Straschewski S, Sinzger C. J Virol. 2012 Jan;86(1):504-12. doi: 10.1128/JVI.05354-11. Epub 2011 Oct 26. | |
2012 | A leucine zipper motif of a tegument protein triggers final envelopment of human cytomegalovirus. | Meissner CS, Suffner S, Schauflinger M, von Einem J, Bogner E. J Virol. 2012 Mar;86(6):3370-82. doi: 10.1128/JVI.06556-11. Epub 2011 Dec 28. |
2012 | Antiviral treatment of cytomegalovirus infection: an update. | Härter G, Michel D. Expert Opin Pharmacother. 2012 Apr;13(5):623-7. doi: 10.1517/14656566.2012.658775. Epub 2012 Feb 2. |
2012 | Cramer J, Mac T, Hogan B, Stauga S, Eberhardt S, Wichmann O, Mertens T, Burchard G. Euro Surveill. 2012 Jan 12;17(2). pii: 20052. | |
2012 | Usmani SM, von Einem J, Frick M, Miklavc P, Mayenburg M, Husmann M, Dietl P, Wittekindt OH. Cell Microbiol. 2012 Mar;14(3):299-315. doi: 10.1111/j.1462-5822.2011.01724.x. Epub 2011 Dec 13. | |
2012 | Rabenau HF, Gottschalk R, Gürtler L, Haberl AE, Hamouda O, Himmelreich H, Korn K, Mertens T, Schmidt KW, Schmiedel S, Spickhoff A, Wirz G, Wutzler P, Wicker S. Bundesgesundheitsblatt Gesundheitsforschung Gesundheitsschutz. 2012 Aug;55(8):937-43. doi: 10.1007/s00103-012-1546-8. German. | |
2012 | Wagner S, Arnold F, Wu Z, Schubert A, Walliser C, Tadagaki K, Jockers R, Mertens T, Michel D. Arch Virol. 2012 May;157(5):935-49. doi: 10.1007/s00705-012-1246-6. Epub 2012 Feb 12. | |
2011 | Schauflinger M, Fischer D, Schreiber A, Chevillotte M, Walther P, Mertens T, von Einem J. J Virol. 2011 Apr;85(8):3821-32. Epub 2011 Feb 2. | |
2011 | M94 is essential for the secondary envelopment of murine cytomegalovirus. | Maninger S, Bosse JB, Lemnitzer F, Pogoda M, Mohr CA, von Einem J, Walther P, Koszinowski UH, Ruzsics Z. J Virol. 2011 Sep;85(18):9254-67. Epub 2011 Jun 29. |
2011 | Jiang XJ, Sampaio KL, Ettischer N, Stierhof YD, Jahn G, Kropff B, Mach M, Sinzger C. Arch Virol. 2011 Dec;156(12):2145-55. Epub 2011 Sep 22. | |
2011 | Schmitt A, Tonn T, Busch DH, Grigoleit GU, Einsele H, Odendahl M, Germeroth L, Ringhoffer M, Ringhoffer S, Wiesneth M, Greiner J, Michel D, Mertens T, Rojewski M, Marx M, von Harsdorf S, Döhner H, Seifried E, Bunjes D, Schmitt M. Transfusion. 2011 Mar;51(3):591-9. doi: 10.1111/j.1537-2995.2010.02940.x. Epub 2010 Dec 6. | |
2011 | von Kries R, Weiss S, Falkenhorst G, Wirth S, Kaiser P, Huppertz HI, Tenenbaum T, Schroten H, Streng A, Liese J, Shai S, Niehues T, Girschick H, Kuscher E, Sauerbrey A, Peters J, Wirsing von König CH, Rückinger S, Hampl W, Michel D, Mertens T. PLoS One. 2011;6(9):e23955. Epub 2011 Sep 7. | |
2011 | Straschewski S, Patrone M, Walther P, Gallina A, Mertens T, Frascaroli G. J Virol. 2011 May;85(10):5150-8. Epub 2011 Mar 2. | |
2011 | Giusti P, Urban BC, Frascaroli G, Albrecht L, Tinti A, Troye-Blomberg M, Varani S. Infect Immun. 2011 Jul;79(7):2727-36. Epub 2011 Apr 4. | |
2011 | Giusti P, Frascaroli G, Tammik C, Gredmark-Russ S, Söderberg-Nauclér C, Varani S. Immunobiology. 2011 Jan-Feb;216(1-2):243-50. Epub 2010 Apr 28. | |
2011 | Reiser M, Wieland A, Plachter B, Mertens T, Greiner J, Schirmbeck R. J Immunol. 2011 Sep 1;187(5):2172-80. Epub 2011 Aug 1. | |
2011 | Telomerase gene mutations are associated with cirrhosis formation. | Hartmann D, Srivastava U, Thaler M, Kleinhans KN, N'kontchou G, Scheffold A, Bauer K, Kratzer RF, Kloos N, Katz SF, Song Z, Begus-Nahrmann Y, Kleger A, von Figura G, Strnad P, Lechel A, Günes C, Potthoff A, Deterding K, Wedemeyer H, Ju Z, Song G, Xiao F, Gillen S, Schrezenmeier H, Mertens T, Ziol M, Friess H, Jarek M, Manns MP, Beaugrand M, Rudolph KL. Hepatology. 2011 May;53(5):1608-17. doi: 10.1002/hep.24217. |